12 Cell-Cell Interaction Analysis
- Understand the concept of cell-cell interactions in spatial transcriptomics
- Learn how to analyze cell-cell interactions using the
CellChatpackage - Interpret the results of cell-cell interaction analysis
- Explore different methods for visualizing cell-cell interactions
12.1 Cell-Cell Interaction Analysis
Cell-cell interactions play a crucial role in various biological processes, including development, immune response, and disease progression. In spatial transcriptomics, analyzing cell-cell interactions can provide insights into how different cell types communicate and influence each other within the tissue microenvironment. We will be using the CellChat package, which provides a comprehensive framework for analyzing and visualizing cell-cell communication networks from single-cell and spatial transcriptomics data.
# Load necessary libraries
library(CellChat)12.2 Prepare Seurat Data for CellChat
First, we need to prepare our Seurat object for CellChat analysis. This involves extracting the expression data and metadata from the Seurat object.
data.input <- visium[["SCT"]]$data
Idents(visium) <- "first_type"
labels <- Idents(visium) #expects the cell labels to be in the identities of the Seurat object
meta <- data.frame(labels = labels, row.names = names(labels)) To use spatial information in the analysis, we can add the spatial coordinates of each spot to the metadata. This step is optional but can provide additional context for interpreting cell-cell interactions. For 10X Visium, this information is in the ‘tissue_positions.csv’ file found in the ‘spatial’ directory, the data stored in the Seurat object is not appropriate for CellChat. We will therefore read in the file directly save the coordinates in a dataframe.
spatial_locs = GetTissueCoordinates(visium, scale = NULL)[ , c("x", "y")] Additionally we need the conversion factor from pixel coordinates to microns. This can be computed as the ratio of the spot diameter in microns (55 microns for 10X Visium) to the spot diameter in pixels (20 pixels for 10X Visium). We need to read the spot diameter in pixels from the ‘scalefactors_json.json’ file found in the ‘spatial’ directory.
scalefactors = jsonlite::fromJSON(txt = file.path("data/mouse_sagittal/spatial", 'scalefactors_json.json'))
spotsize = 65 # the theoretical spot size (um) in 10X Visium
conversion_factor = spotsize/scalefactors$spot_diameter_fullres
spatial_factors = data.frame(ratio = conversion_factor, tol = spotsize/2)Now we can check the distances of the spots to each other, which will be used in the CellChat analysis. For 10X Visium v1 data it should be roughly 100 microns between the centers of two spots.
#check distances of the spots to each other
dist <- computeCellDistance(coordinates = spatial_locs, ratio = spatial_factors$ratio, tol = spatial_factors$tol)
min(dist[dist!=0]) 12.3 Create CellChat Object & Set the ligand-receptor database
Next, we create a CellChat object using the expression data and metadata as well as the spatial coordinates we have just prepared.
cellchat <- createCellChat(object = data.input, meta = meta, group.by = "labels", datatype = "spatial", coordinates = spatial_locs, spatial.factors = spatial_factors)Then we load and set the ligand-receptor interaction database. CellChat provides several built-in databases, and we will use the mouse database for this example. We are excluding the “Non-protein Signaling” category for our analysis, which is a standard subset of interactions to consider. We will also remove unused expression data from the CellChat object to speed up the analysis, matching the selected database subset.
CellChatDB <- CellChatDB.mouse
showDatabaseCategory(CellChatDB)
# use all CellChatDB except for "Non-protein Signaling" for cell-cell communication analysis
CellChatDB.use <- subsetDB(CellChatDB)
# set the used database in the object
cellchat@DB <- CellChatDB.use
# subset the expression data of signaling genes for saving computation cost
cellchat <- subsetData(cellchat) 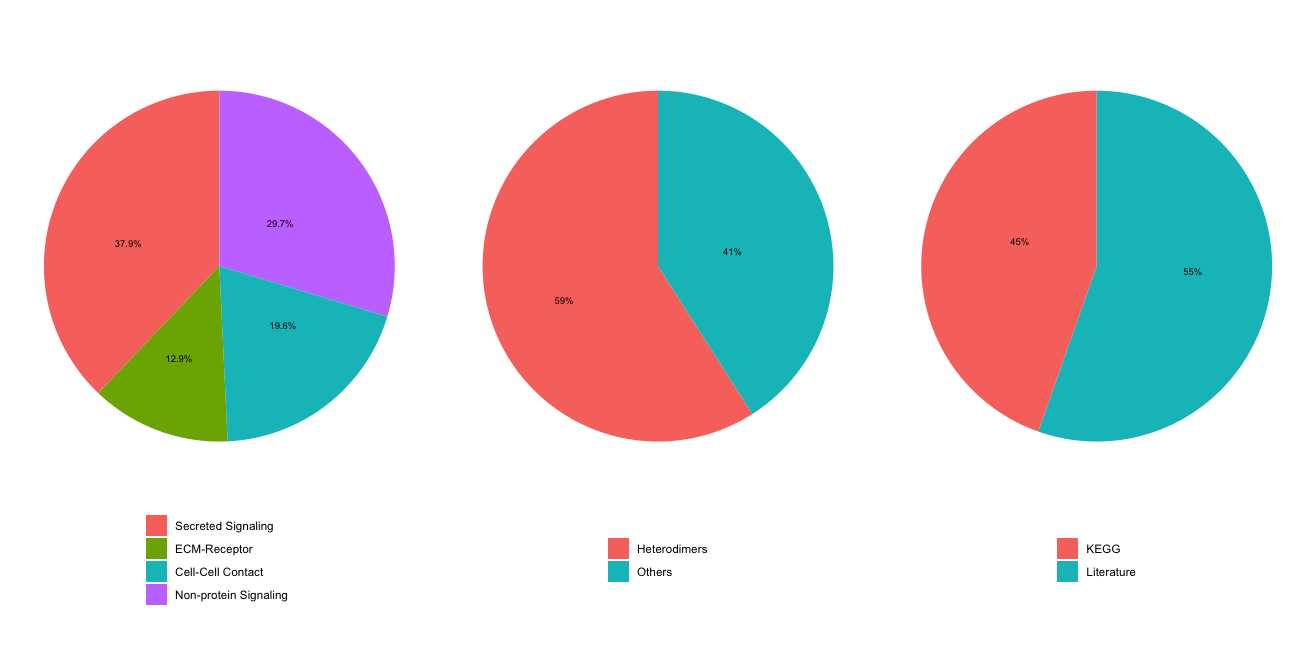
These are the categories of ligand-receptor interactions available in the CellChat mouse database. Each category represents a different type of signaling mechanism that can occur between cells.
12.4 Identify Overexpressed Genes and Interactions
We will identify overexpressed genes and interactions in the dataset. This step helps to focus the analysis on the most relevant genes and interactions. CellChat offers parallel processing, but this often actually causes more issues than speedup, so we will not be using it. To compute the communication probability at the signaling gene level, we will use the “triMean” method, which is a robust method for estimating communication probabilities. We need to set the ‘contact.range’ parameter to define the maximum distance for considering cell-cell interactions. Here, we set it to 100 microns, which is the expected distance between the centers of two spots in 10X Visium data. We will also filter the interactions to only consider those that are supported by at least 10 cells.
# Identify overexpressed genes and interactions
cellchat <- identifyOverExpressedGenes(cellchat)
cellchat <- identifyOverExpressedInteractions(cellchat)
#The following function computes the communication probability at the signaling pathway level
#If it takes a long time to compute, so you can skip it and load precomputed results instead
cellchat <- computeCommunProb(cellchat, type = "triMean", contact.range = 100, trim = 0.1)
cellchat <- computeCommunProbPathway(cellchat)
cellchat <- filterCommunication(cellchat, min.cells = 10) #only consider interactions that are supported by at least 10 cells
cellchat <- aggregateNet(cellchat)
# cellchat <- loadRDS("precomputed/mouse_brain_cellchat.rds") #load precomputed resultsThe number of highly variable ligand-receptor pairs (output of the identifyOverExpressedInteractions function) identified for this dataset is 1467.
We have now computed the cell-cell communication probabilities and aggregated the communication network at the cell type level. The resulting CellChat object contains information about the identified interactions, their probabilities, and the overall communication network between different cell types in the dataset.
12.5 Visualize Cell-Cell Communication Network
Finally, we can visualize the cell-cell communication network using various plotting functions provided by the CellChat package. We will create a circle plot to visualize the overall communication network and a heatmap to show the interaction strength between different cell types as well as a bar plot ranking the signaling pathways by their acttivity scores. We will both choose to visualise two specific pathways of interest: “COLLAGEN” and “LAMININ” together, as well as “LAMININ” alone.
Laminin is a key component of the extracellular matrix and plays a crucial role in cell adhesion, differentiation, migration, and signaling. It is particularly important in the context of neural development and function, making it a relevant pathway to investigate in brain tissue.
Collagen is another major component of the extracellular matrix and is involved in providing structural support to tissues. It also plays a role in cell adhesion and signaling, making it another important pathway to consider in the context of cell-cell interactions.
#find all significant signalling pathways
pathways.show.all <- cellchat@netP$pathways
pathways.selected <- c("COLLAGEN", "LAMININ")
laminin <- c("LAMININ")
# Circle plos of the overall communication network
groupSize <- as.numeric(table(cellchat@idents))
netVisual_circle(cellchat@net$count, vertex.weight = groupSize, weight.scale = T , label.edge= F, title.name = "Number of interactions", top = 0.3)
netVisual_circle(cellchat@net$weight, vertex.weight = groupSize, weight.scale = T , label.edge= F, title.name = "Interaction weights/strength", top = 0.3) The first circle plot shows the number of interactions between different cell types, while the second plot shows the interaction weights or strengths. The size of each node represents the number of cells in that cell type, and the thickness of the edges represents the strength of the interactions between cell types.
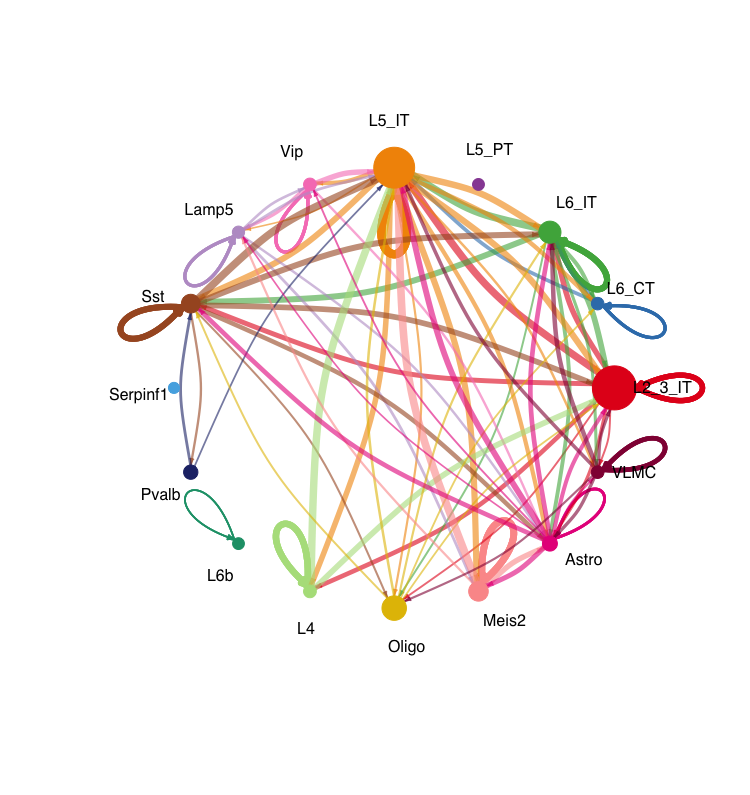
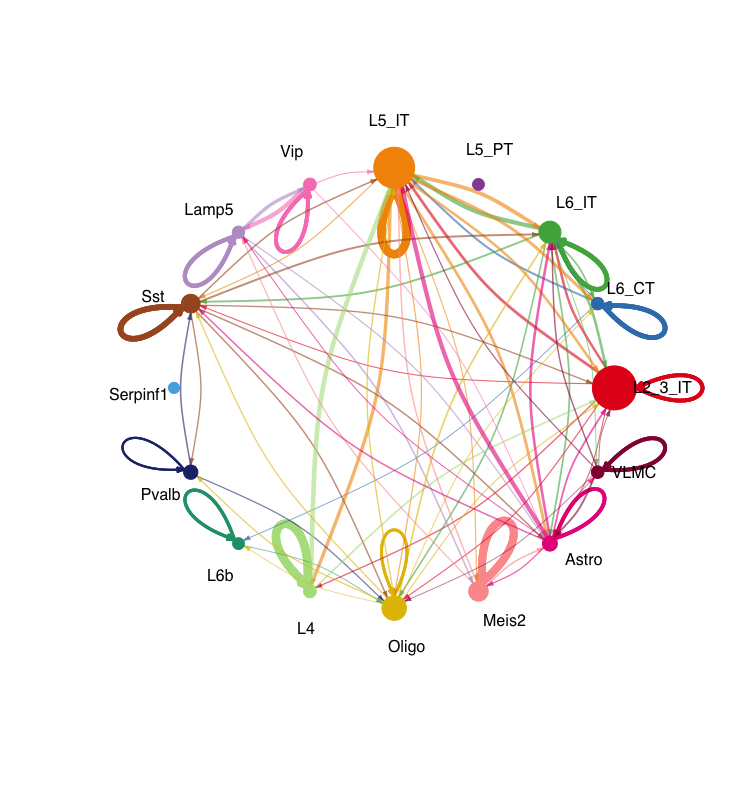
# Heatmap of interaction strength between cell types
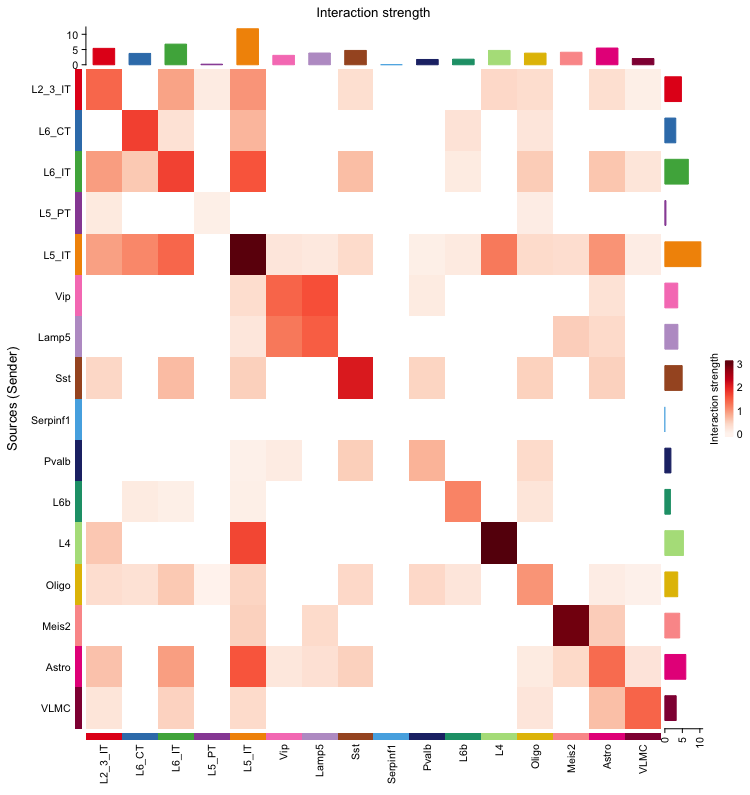
netVisual_heatmap(cellchat, measure = "weight") The heatmap visualizes the interaction strength between different cell types. The color intensity represents the strength of the interactions, with darker colors indicating stronger interactions. This heatmap provides a comprehensive overview of the communication network, allowing us to identify which cell types are more actively communicating with each other.
#Bar plot of the contribution of each ligand-receptor interaction pair to the selected pathways
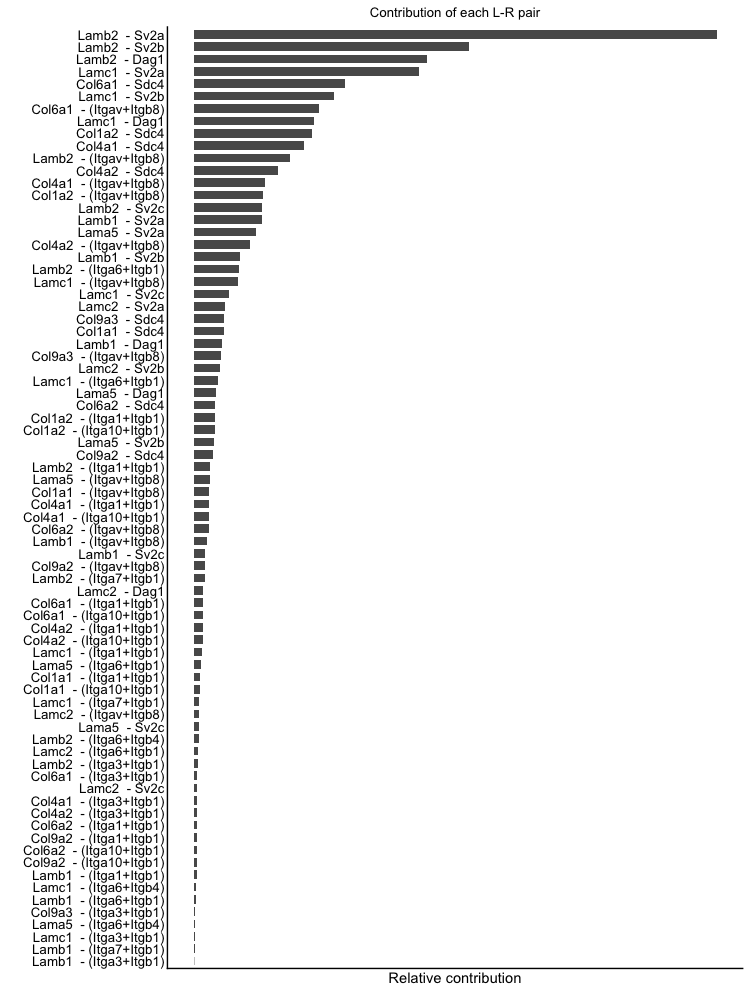
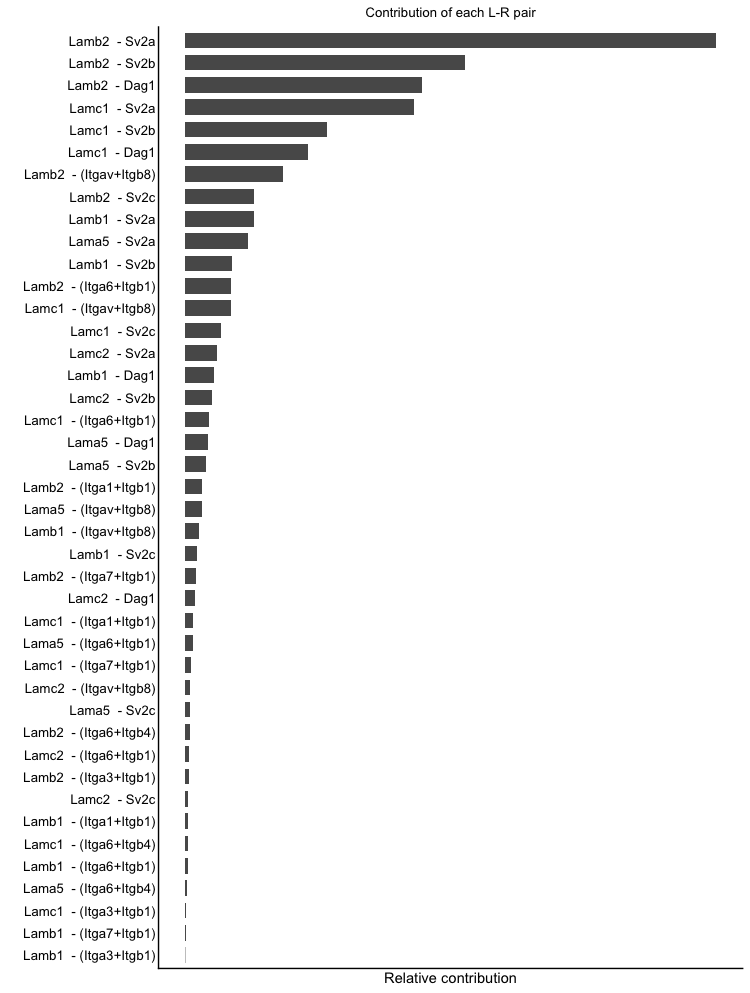
netAnalysis_contribution(cellchat, signaling = pathways.selected)
netAnalysis_contribution(cellchat, signaling = laminin)The bar plots show the contribution of each ligand-receptor interaction pair to the selected pathways. The height of each bar represents the contribution score, which indicates how much each interaction pair contributes to the overall signaling activity of the pathway. This visualization helps to identify the key ligand-receptor pairs that are driving the signaling activity in the selected pathways.
# Bubble Plot of ligand-receptor interactions from one celltype (the 1st, L2/3 IT) to other celltypes (the 3rd to 5th)
netVisual_bubble(cellchat, sources.use = 1, targets.use = c(3:5), remove.isolate = FALSE)The bubble plot visualizes the ligand-receptor interactions from one cell type (L2/3 IT) to other cell types (the 3rd to 5th). Each bubble represents a ligand-receptor pair, with the size of the bubble indicating the strength of the interaction. This plot allows us to see how a specific cell type communicates with other cell types in the tissue.

#Centrality scores of the selected pathways
cellchat <- netAnalysis_computeCentrality(cellchat, slot.name = "netP")
netAnalysis_signalingRole_network(cellchat, signaling = pathways.selected, width =10, height = 4)
# Incoming vs outgoing signalling strength for all signalling roles in the network per celltype for all pathways and separately for the laminin pathway
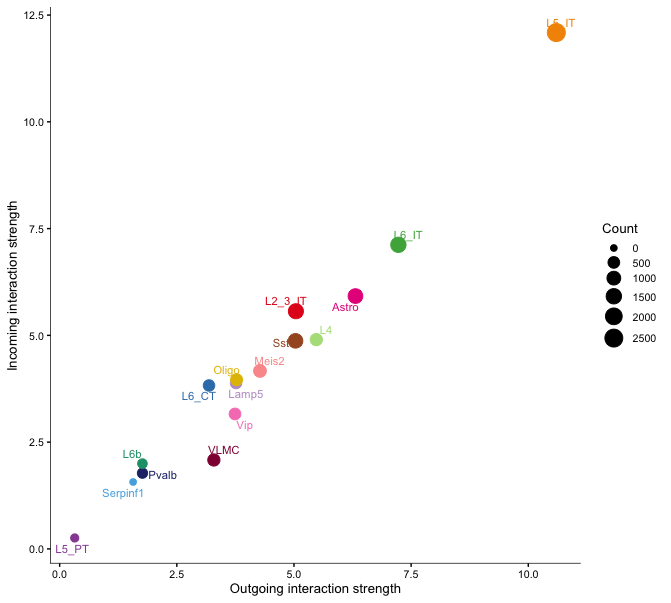
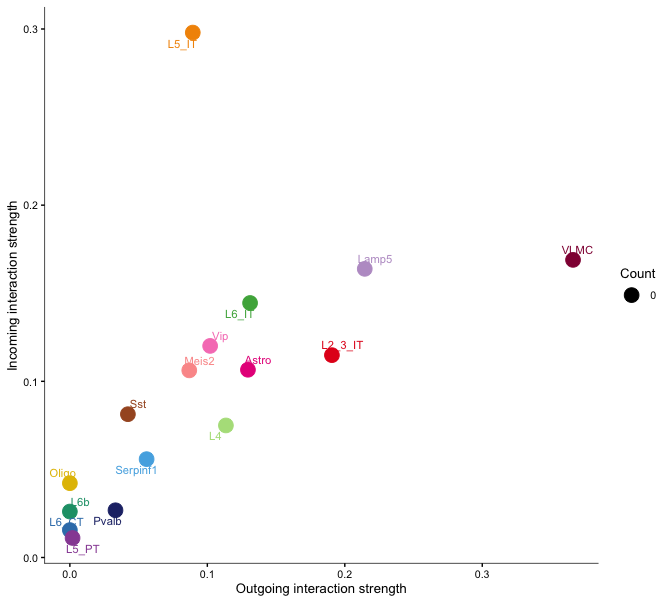
netAnalysis_signalingRole_scatter(cellchat)
netAnalysis_signalingRole_scatter(cellchat, signaling = c("LAMININ"))The first plot shows the centrality scores of the selected pathways, indicating the importance of each cell type in the communication network. 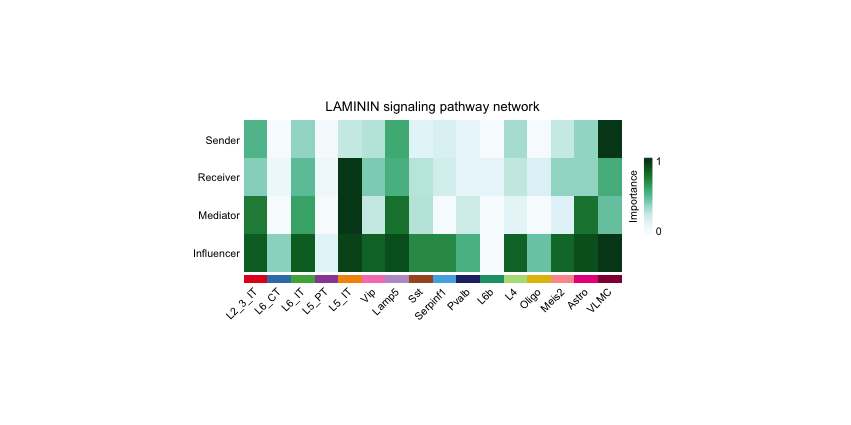
The second and third plots show the incoming vs outgoing signaling strength for all signaling roles in the network per cell type for all pathways and separately for the laminin pathway. These scatter plots help to identify which cell types are primarily sending or receiving signals in the network.
Of course, we can also visualize specific signaling pathways of interest in spatial context.
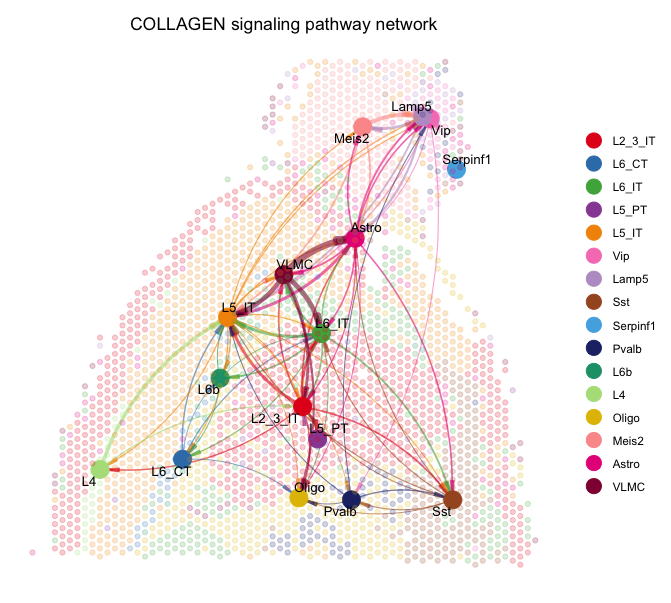
netVisual_aggregate(cellchat, signaling = pathways.selected, layout = "spatial", edge.width.max = 2, vertex.size.max = 1, alpha.image = 0.2, vertex.label.cex = 3.5)
netVisual_aggregate(cellchat, signaling = laminin, layout = "spatial", edge.width.max = 2, vertex.size.max = 1, alpha.image = 0.2, vertex.label.cex = 3.5)The spatial plots visualize the selected signaling pathways in the context of the tissue. The nodes represent different cell types, and the edges represent the interactions between them. The layout is based on the spatial coordinates of the spots, allowing us to see how cell-cell interactions are distributed across the tissue.
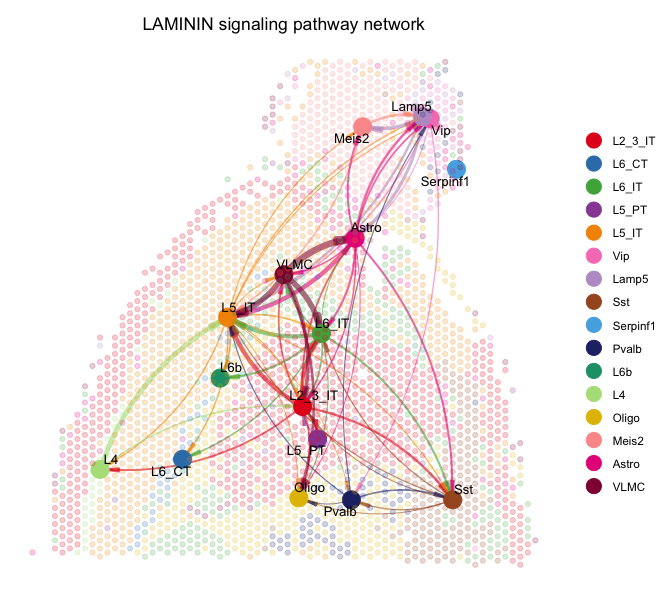
There is a plethora of other visualization and investigation options available in the CellChat package. You can explore these options in the CellChat documentation to find the best way to visualize your specific data and research questions.
12.6 Conclusion
In this section, we have explored how to analyze cell-cell interactions using the CellChat package in R. We prepared the Seurat object for CellChat analysis, identified overexpressed genes and interactions, and visualized the cell-cell communication network. Understanding cell-cell interactions can provide valuable insights into tissue architecture and function, helping to unravel complex biological processes.
12.7 Summary
- Cell-cell interactions are crucial for understanding tissue microenvironments and biological processes.
- The
CellChatpackage provides a comprehensive framework for analyzing and visualizing cell-cell communication networks. - Preparing the Seurat object and identifying overexpressed genes and interactions are essential steps in the analysis.
- For spatial transcriptomics data, incorporating spatial coordinates can enhance the analysis of cell-cell interactions.
- Visualization techniques such as circle plots, heatmaps, and bar plots help interpret the results of cell-cell interaction analysis.