7 Clustering of Spatial Transcriptomics Data
- Cluster spatial transcriptomics data using Seurat
- Visualize clusters using UMAP and spatial plots
- Identify marker genes for clusters
- Understand the interpretation of clustering results
7.1 Clustering Spatial Transcriptomics Data
Clustering is a crucial step in spatial transcriptomics analysis, allowing us to group cells based on their gene expression profiles. In Seurat, basic clustering can be performed using the FindClusters function after dimensionality reduction techniques like PCA or UMAP. This clustering helps to identify distinct cell populations, but is not spatially aware. Seurat offers different clustering algorithms, with the Louvain algorithm being the default choice. The resolution parameter in the FindClusters function controls the granularity of the clustering, with higher values leading to more clusters. The default resolution is 0.8, but this can be adjusted based on the dataset and the desired level of detail. As default, these clusters are stored in the metadata of the Seurat object under the name “seurat_clusters”, which can be changed using the cluster.name parameter.
Before clustering we need to compute a nearest neighbor graph using the FindNeighbors function. This function constructs a graph based on the PCA results, which is then used for clustering.
#load dplyr for data manipulation
library(dplyr)
#Perform default clustering using Louvain algorithm on the PCA results
visium <- FindNeighbors(visium, reduction = "pca", dims = 1:30)
visium <- FindClusters(visium, resolution = 0.5, cluster.name = "Louvain_05") We can visualize the clustering results using UMAP or spatial plots. The DimPlot function can be used to create UMAP plots, while the SpatialDimPlot function is used for spatial visualizations.
#UMAP plot of clusters
umap_louvain05 <- DimPlot(visium, reduction = "umap", group.by = "Louvain_05", label = TRUE) + ggtitle("Louvain Clusters (res=0.5)")
#Spatial plot of clusters
spatial_louvain05 <- SpatialDimPlot(visium, group.by = "Louvain_05", label = TRUE) + ggtitle("Louvain Clusters (res=0.5)") We would like to compare these results to the ones achieved with Leiden clustering.
#Perform Leiden clustering
visium <- FindClusters(visium, resolution = 0.5, algorithm = 4, cluster.name = "Leiden_05")
#UMAP plot of Leiden clusters
umap_leiden05 <- DimPlot(visium, reduction = "umap", group.by = "Leiden_05", label = TRUE) + ggtitle("Leiden Clusters (res=0.5)")
#Spatial plot of Leiden clusters
spatial_leiden05 <- SpatialDimPlot(visium, group.by = "Leiden_05", label = TRUE) + ggtitle("Leiden Clusters (res=0.5)")
umap_louvain05 + umap_leiden05
spatial_louvain05 + spatial_leiden05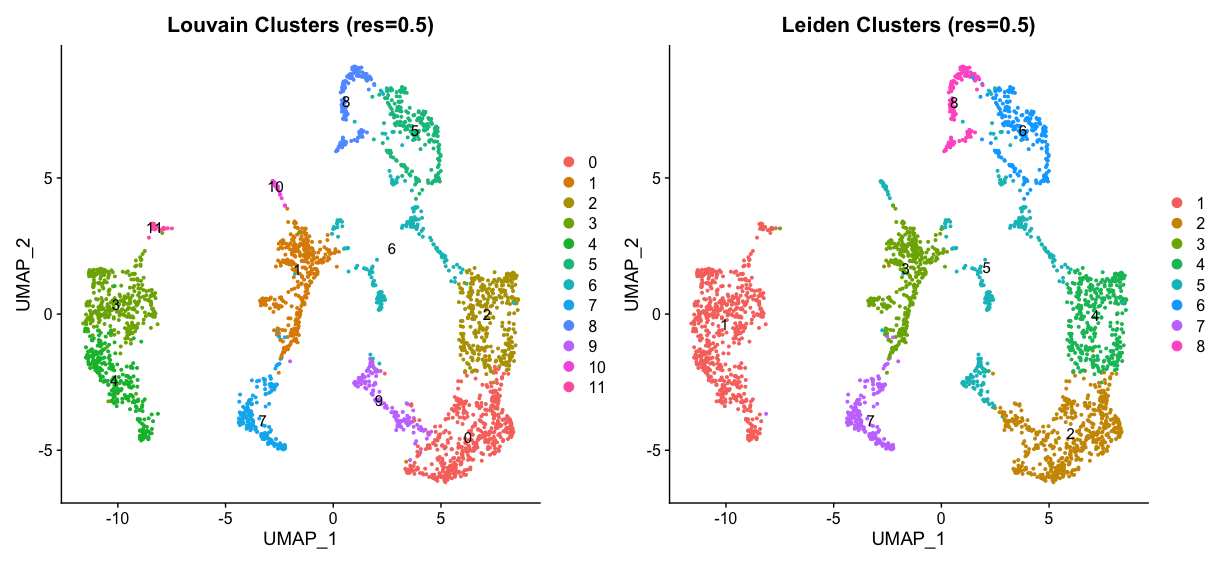
It’s hard to decide which clustering is better just by looking at the plots. Currently it looks like we have not chosen the optimal resolution, as we have big clusters that could be further subdivided. We can try a higher resolution to see if we can identify more distinct clusters.
We can look at the spatial distribution of the clusters to see if they make sense biologically. If we see clusters that are spatially coherent and correspond to known tissue structures, it suggests that the clustering is capturing meaningful biological variation.
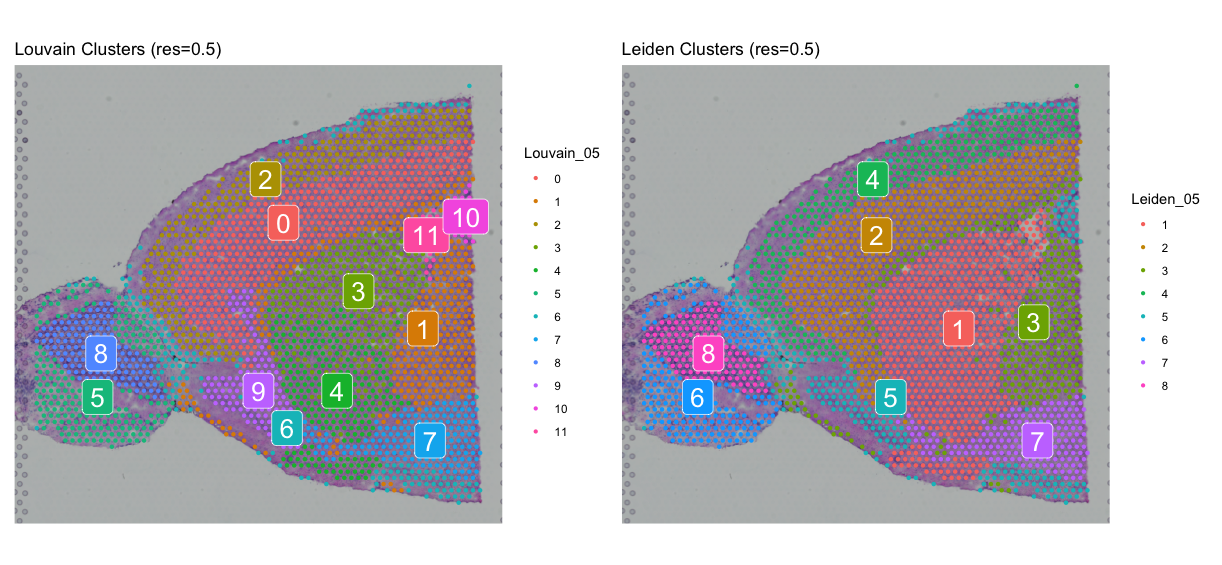
Now, let’s try a higher resolution of 0.8 for both clustering algorithms and compare the results.
#Perform Louvain clustering with higher resolution
visium <- FindClusters(visium, resolution = 0.8, cluster.name = "Louvain_08")
#UMAP plot of clusters
umap_louvain08 <- DimPlot(visium, reduction = "umap", group.by = "Louvain_08", label = TRUE) + ggtitle("Louvain Clusters (res=0.8)")
#Spatial plot of clusters
spatial_louvain08 <- SpatialDimPlot(visium, group.by = "Louvain_08", label = TRUE) + ggtitle("Louvain Clusters (res=0.8)")
#Perform Leiden clustering with higher resolution
visium <- FindClusters(visium, resolution = 0.8, algorithm = 4, cluster.name = "Leiden_08")
#UMAP plot of Leiden clusters
umap_leiden08 <- DimPlot(visium, reduction = "umap", group.by = "Leiden_08", label = TRUE) + ggtitle("Leiden Clusters (res=0.8)")
#Spatial plot of Leiden clusters
spatial_leiden08 <- SpatialDimPlot(visium, group.by = "Leiden_08", label = TRUE) + ggtitle("Leiden Clusters (res=0.8)")
umap_louvain08 + umap_leiden08
spatial_louvain08 + spatial_leiden08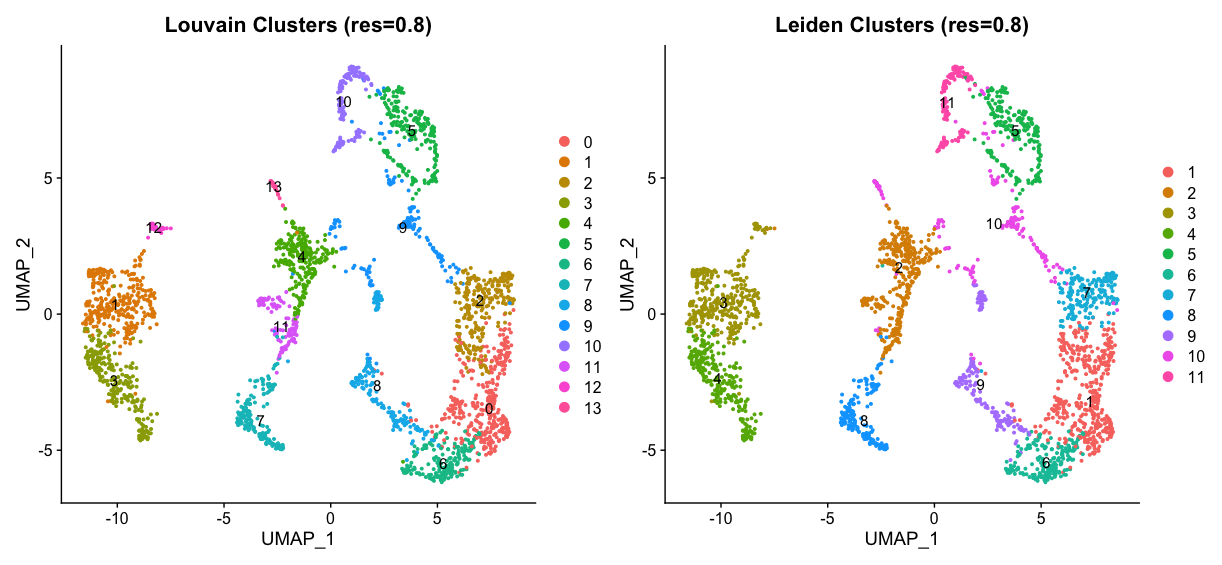
Clearly Louvain clustering is more sensitive to changes in the resolution parameter, as we can see a big difference between resolution 0.5 and 0.8. The Leiden algorithm is more robust to changes in the resolution parameter, as the clustering results are more similar between the two resolutions. However, the Leiden clustering at resolution 0.8 seems to be more reasonable than the Louvain clustering at the same resolution, as we can see more distinct clusters in the spatial plot.
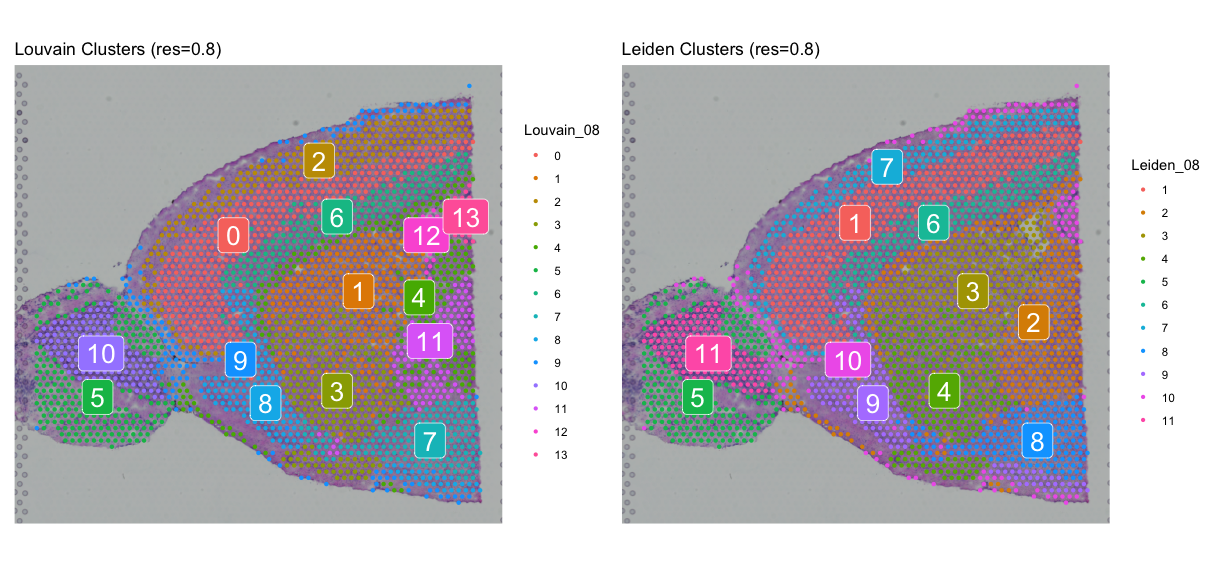
Apart from visual inspection, we can also use domain knowledge to evaluate the clustering results. For example, if we expect 20 cell types in the tissue, we can choose the clustering that results in a similar number of clusters. We have to remember though, that clusters in spatial transcriptomics do not necessarily correspond to cell types, as there can be multiple clusters per cell type due to different states or subtypes.
This type of clustering is not spatially aware, meaning that it does not take into account the spatial location of the spots. There are more advanced clustering methods that do consider spatial information, such as BANKSY, which we will cover later in this course.
7.1.1 Identifying Marker Genes for Clusters
After clustering, we can identify marker genes for each cluster using the FindAllMarkers function. This function performs differential expression analysis to find genes that are significantly upregulated in each cluster compared to all other clusters. We are going to select only positive markers, meaning genes that are more highly expressed in the cluster of interest compared to other clusters. We will also set a minimum percentage of cells expressing the gene (min.pct) and a log-fold change threshold (logfc.threshold) to filter the results.
#Set the identity class to the clustering we want to find markers for
Idents(visium) <- visium$Leiden_08
# Identify marker genes for each cluster
markers <- FindAllMarkers(visium, only.pos = TRUE, min.pct = 0.25, logfc.threshold = 0.25, pvalue.cutoff = 0.05)
# View top markers for each cluster
top10 <- markers %>% group_by(cluster) %>% top_n(n = 10, wt = avg_log2FC)
#Visualise the 4 top markers for cluster 1
fp <- SpatialFeaturePlot(visium, features = top10[top10$cluster==1,]$gene[1:4], ncol = 2)
# Visualise the spatial location of cluster 1
sp <- SpatialDimPlot(visium, cells.highlight = WhichCells(visium, idents = "1")) + ggtitle("Cluster 1 Spatial Location")
fp + sp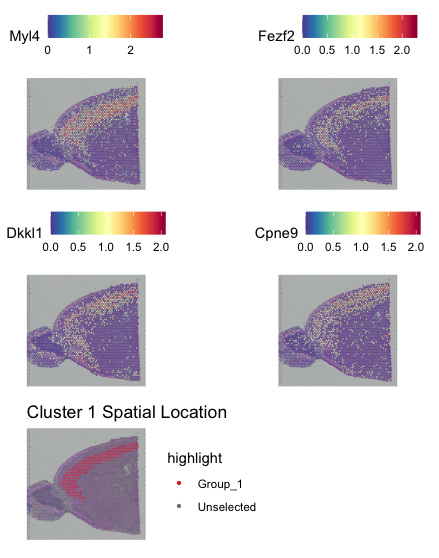
By default, the FindAllMarkers function uses the Wilcoxon rank-sum test for differential expression analysis. Other tests can be specified using the test.use parameter. The results include the average log-fold change, p-value, and adjusted p-value for each gene in each cluster. Another option is to use the MAST test, which is specifically designed for single-cell data and can account for cellular detection rates, but is computationally more intensive and not always optimal for spatial transcriptomics data.
7.2 Conclusion
In this chapter, we have learned how to perform clustering on spatial transcriptomics data using Seurat. We have explored different clustering algorithms, such as Louvain and Leiden, and discussed the impact of the resolution parameter on the clustering results. We have also identified marker genes for each cluster using differential expression analysis. Clustering is a crucial step in spatial transcriptomics analysis, as it helps to identify distinct cell populations and understand the underlying biology of the tissue.
7.3 Summary
- Clustering is essential for identifying distinct cell populations in spatial transcriptomics data.
- Seurat provides multiple clustering algorithms, with Louvain and Leiden being popular choices.
- The resolution parameter controls the granularity of the clustering, with higher values leading to more clusters.
- Marker genes for each cluster can be identified using the
FindAllMarkersfunction, which performs differential expression analysis.