9 Publication-quality visualisations
- Create clear and informative molecular visualisations using ChimeraX.
- Combine structural representations such as cartoons, surfaces, and atoms.
- Adjust lighting, background, and outlines to improve figure clarity.
- Generate rotating or animated movies of molecular structures.
- Use tiled views to compare multiple structures simultaneously.
High-quality molecular visualisations are an important part of communicating structural biology results. Figures used in publications or presentations should aim to be:
- informative - highlighting the structural features relevant to the question
- clear - avoiding unnecessary clutter
- visually consistent - using colour and representation carefully
ChimeraX provides a wide range of tools for producing such figures.
The ChimeraX gallery showcases many examples of publication-quality visualisations. Each example includes a downloadable .cxc script that reproduces the image using ChimeraX commands.
Studying these scripts is often a useful way to learn new visualisation techniques.
9.1 Surface representation
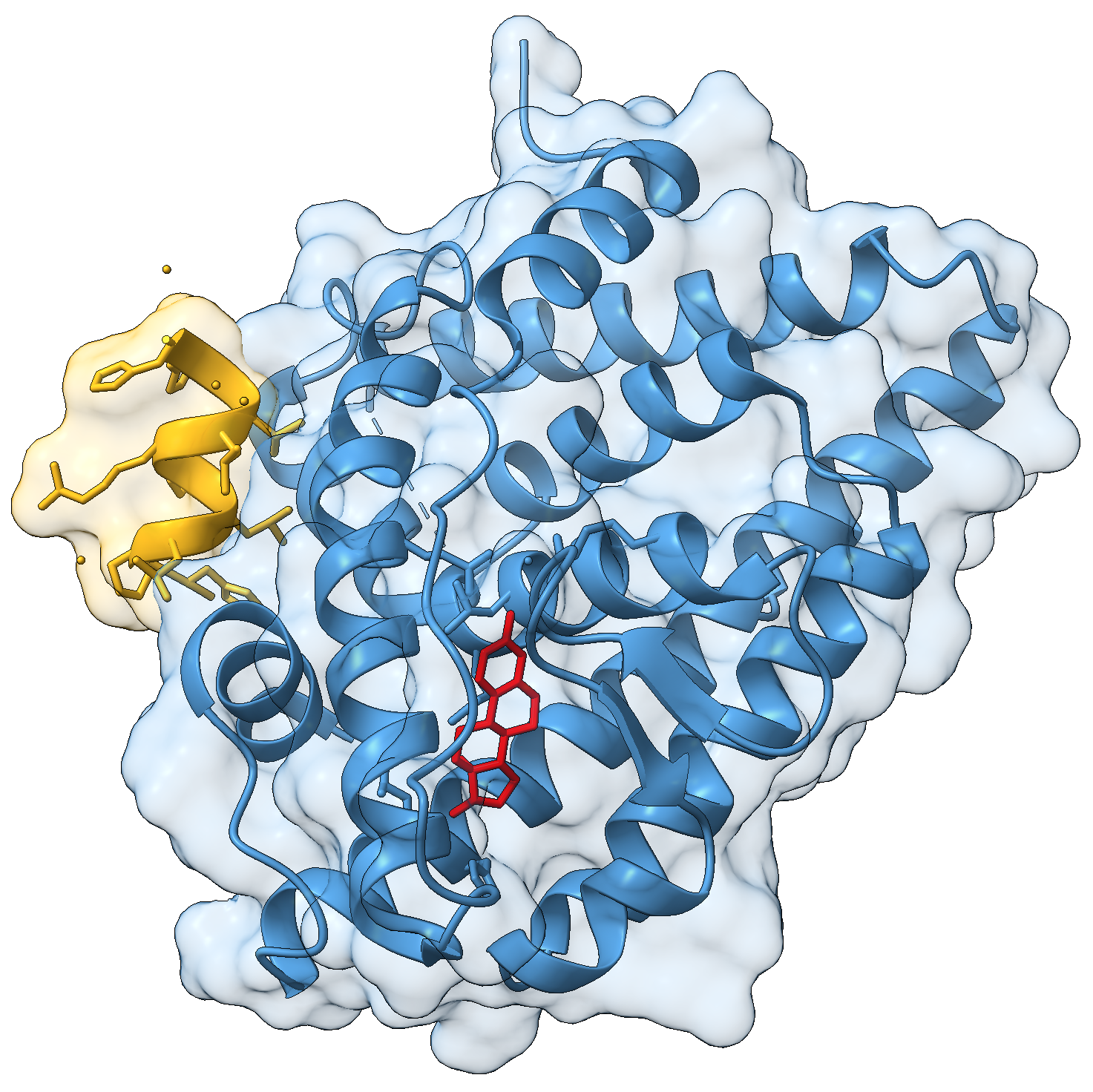
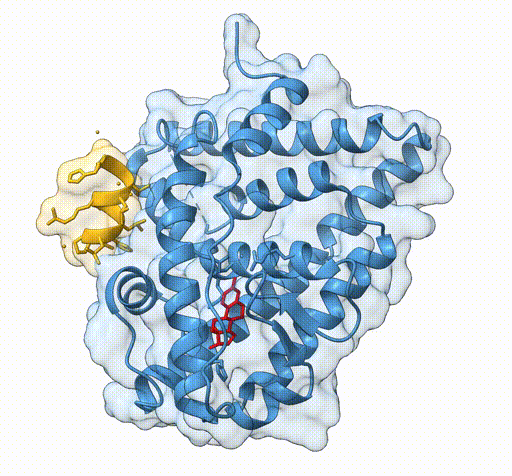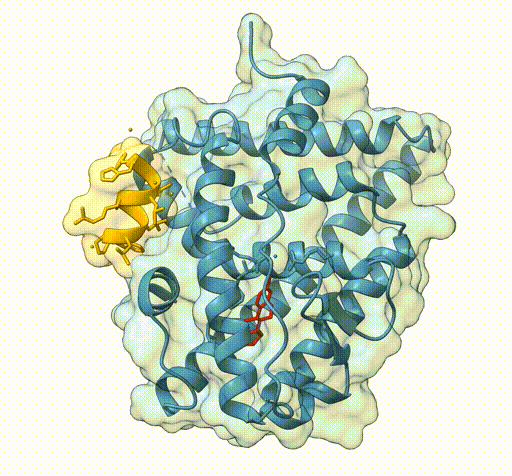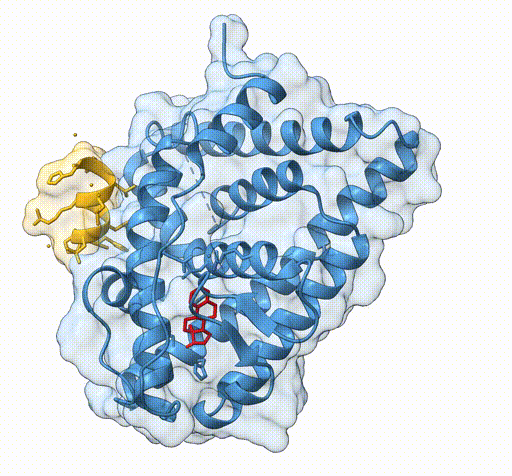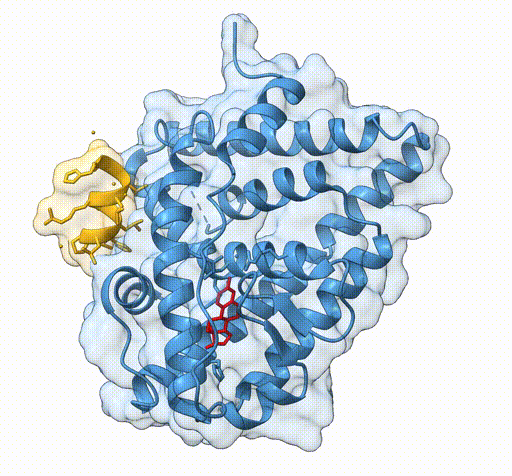
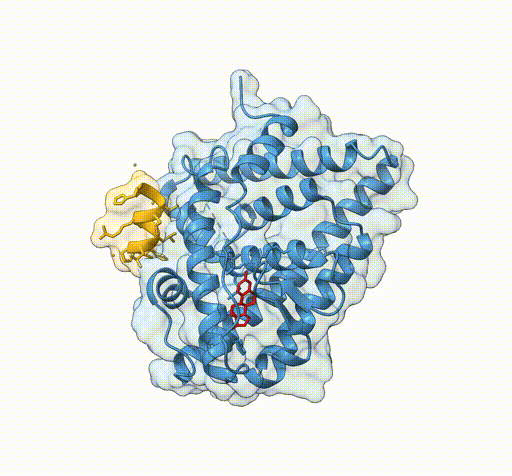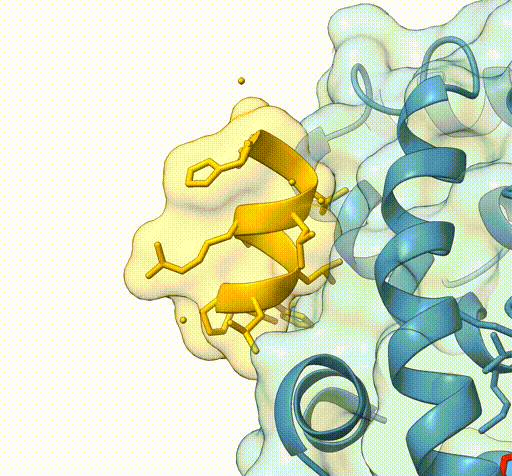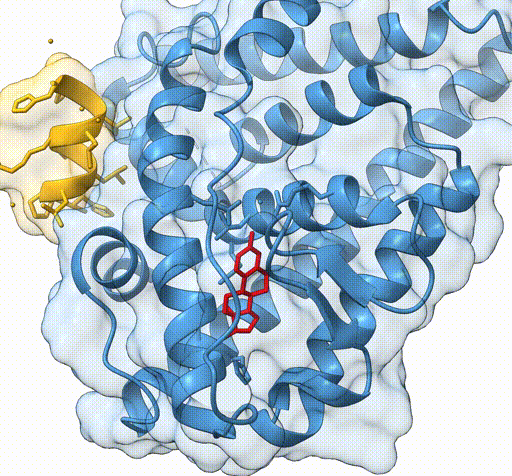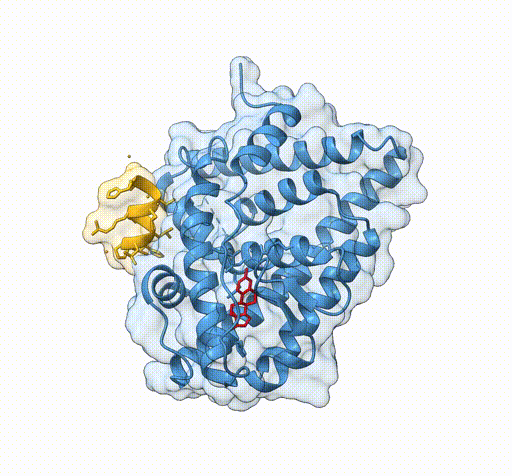
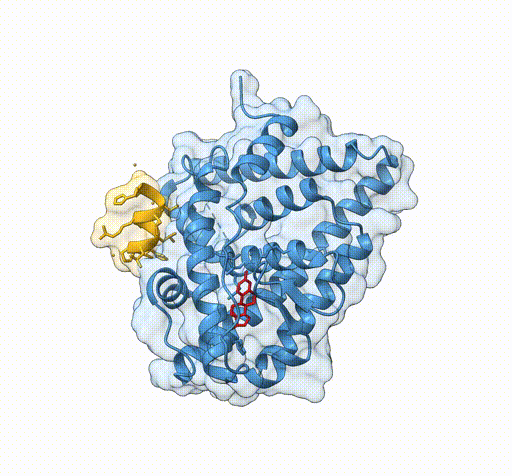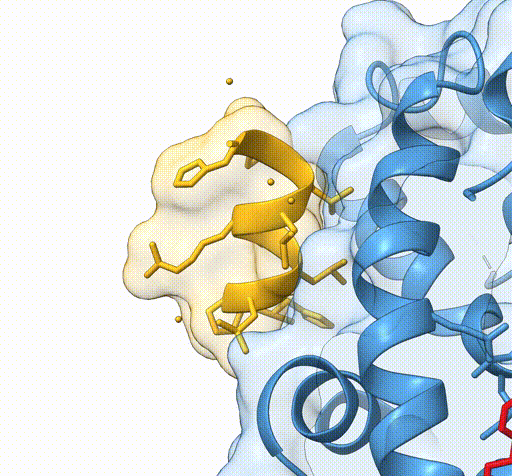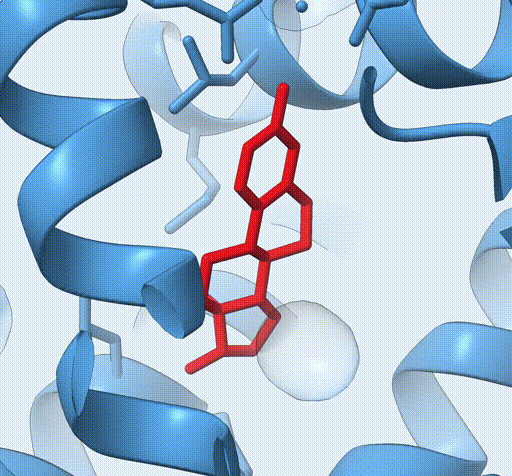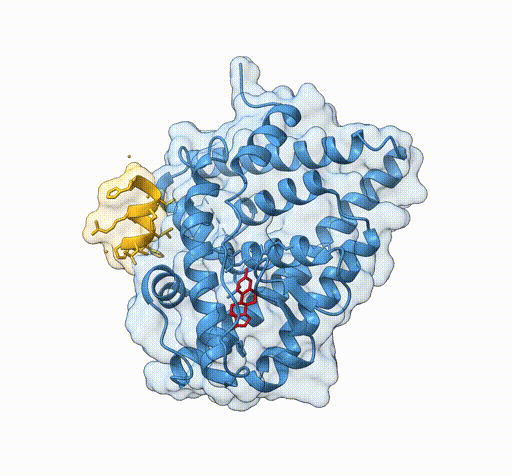
Consider the following view of the estrogen receptor in complex with a conserved peptide motif characteristic of activator proteins (PDB: 4J26).
This visualisation combines several features to highlight the interaction between the receptor and the peptide.
Below are the commands used to generate the figure:
close
open 4J26
delete /A,I
set bgcolor white
label delete
light simple
graphics silhouettes true width 1
show /J atoms
color /B steelblue
color /J goldenrod
color :EST red
surface /B transparency 85
surface /J transparency 85Let’s break these down step-by-step.
We begin by loading the structure and simplifying the model.
The crystal structure contains two receptor-peptide complexes, so we remove chains A and I to keep only one complex.
delete /A,INext we adjust the global visual settings.
set bgcolor white
label delete
light simple
graphics silhouettes true width 1These commands:
- set a white background, which is standard for publications
- remove default residue labels
- use simple lighting to improve contrast
- add silhouettes (black outlines) around objects
Silhouettes are particularly useful because they help the structure stand out clearly against the background.
We then highlight the relevant structural components.
show /J atoms
color /B steelblue
color /J goldenrod
color :EST redHere we:
- display the coactivator peptide atoms
- colour the receptor chain blue
- colour the peptide gold
- colour the ligand red
Using distinct colours makes it easier to identify the different molecular components.
Finally, we display transparent molecular surfaces for both chains:
surface /B transparency 85
surface /J transparency 85The transparent surface illustrates the shape of the binding interface, while still allowing the underlying secondary structure to remain visible.
9.2 Creating a movie
Movies can help illustrate structural features that are difficult to appreciate in a static image.
ChimeraX supports several types of automatic motion:
rock- oscillating back-and-forth motionroll- continuous 360 degree rotationwobble- figure-eight style motionturn- flexible rotation around arbitrary axes
The general workflow for creating a movie is:
- start recording
- perform the desired movement
- stop recording
- encode the frames into a video file
9.2.1 Example: rocking animation
We can create a rocking animation of the estrogen receptor using:
rockTo stop the motion:
stopTo record a movie, we use the movie record command. It is helpful to record complete motion cycles, so that the movie loops smoothly.
According to the ChimeraX documentation, the default rocking motion completes one cycle in 136 frames. We therefore record one full cycle using:
movie record
rock
wait 136
stopFinally, we encode the movie as an MP4 file:
movie encode 4j26_rock.mp4 framerate 60The framerate option determines how many frames are shown per second in the final movie.
9.3 Animated views
ChimeraX also allows you to define named views and smoothly transition between them. This approach is useful for guiding the viewer through different parts of a structure.
First we define a set of views:
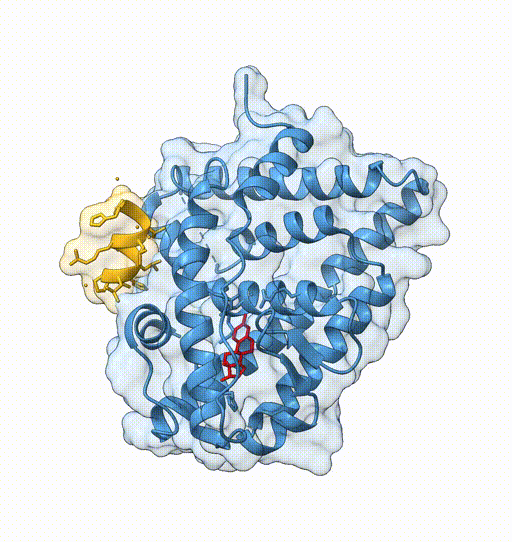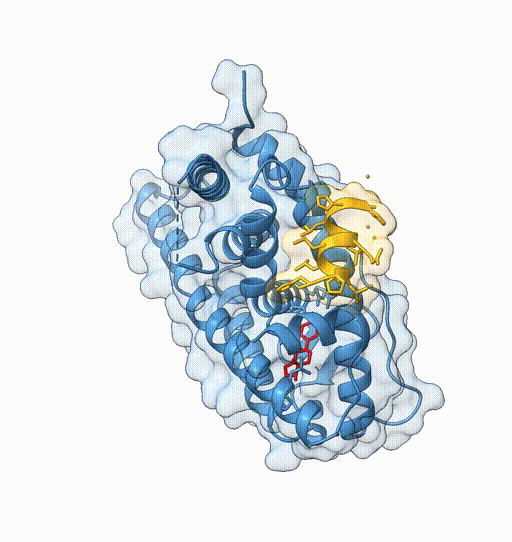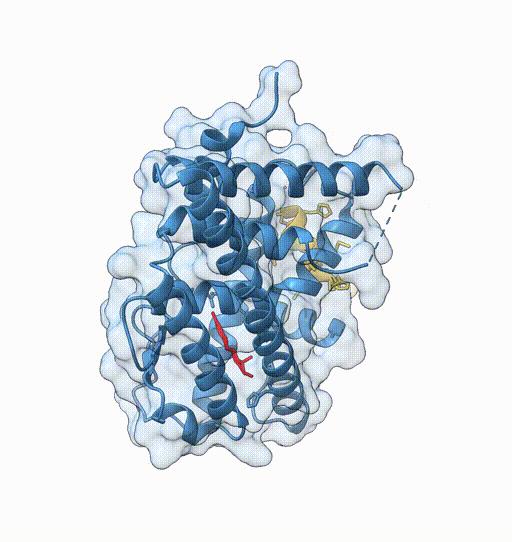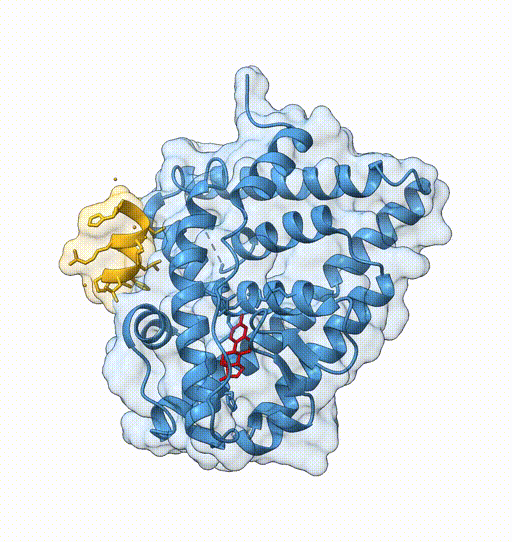
- View 1: entire protein
- View 2: coactivator peptide
- View 3: ligand binding site
view protein
view name 1
view /J
view name 2
view ligand
view name 3Each view name command stores the current camera position. We can then transition between these views while recording a movie. The syntax:
view <name> <frames>controls how long the transition takes.
For example:
view 1
movie record
view 2 60
wait 80
view 1 30
wait 30
view 3 60
wait 80
view 1 30
wait 30
movie encode 4j26_views.mp4 framerate 30Deciding on the framerates for each view and transition can require some trial and error, but it helps to think about it in relation to the framerate used to encode the movie. In this example:
- the movie is encoded at 30 frames per second
- a transition of 60 frames corresponds to 2 seconds
- pauses are introduced using the
waitcommand
9.4 UniProt annotations
Functional annotations can also be incorporated into visualisations.
For example, we can retrieve UniProt information for the estrogen receptor. The UniProt identifier for the receptor is P03372:
open P03372 from uniprot format uniprotThis opens the sequence annotation panel, where features such as domains, mutations, and binding sites can be inspected. Selecting a feature highlights the corresponding residues in the structure.
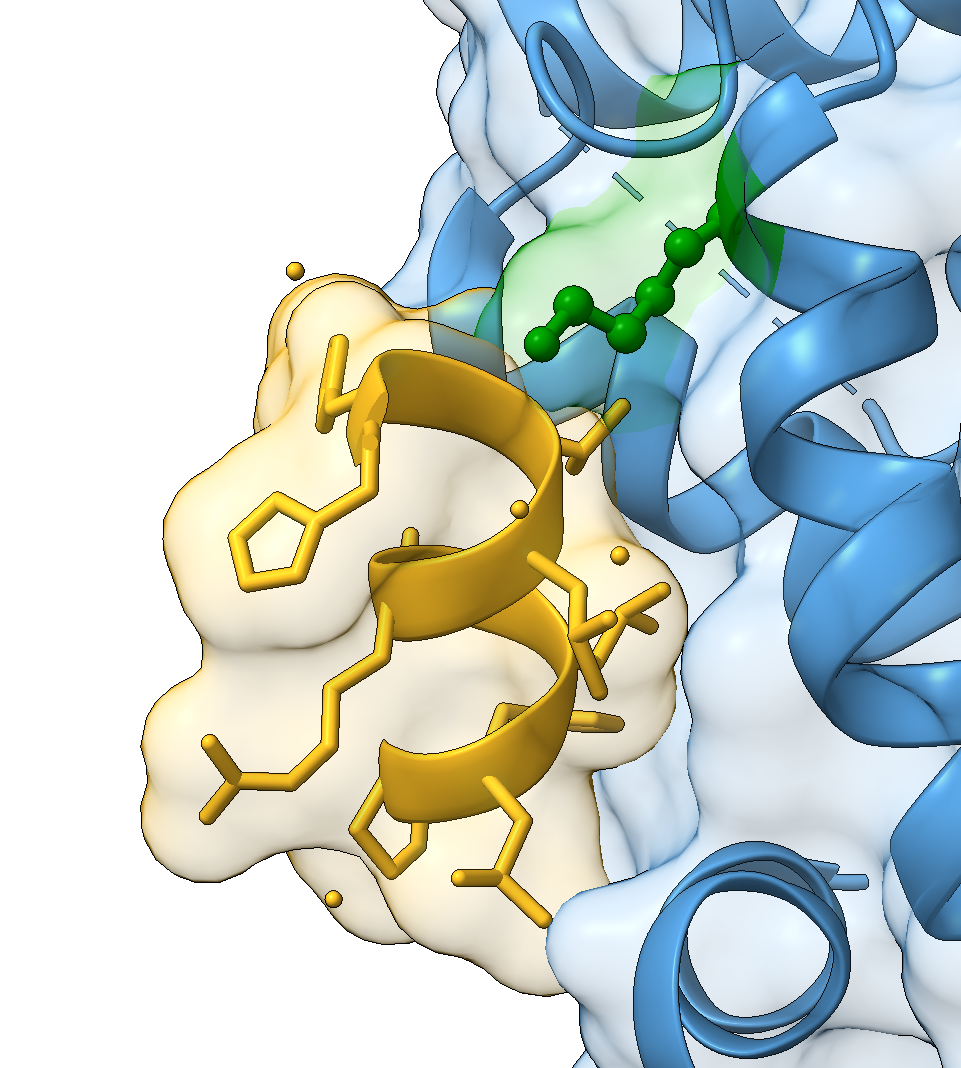
For example, one of the mutations annotated in UniProt is described as: “K → R completely inactive in positive regulation of DNA-binding transcription factor activity”. Once selected, the residue can be highlighted in the structure:
select /B:314
show sel atoms
style sel ball
colour sel green
select clearHighlighting biologically important residues is often helpful when preparing figures that illustrate functional mechanisms.
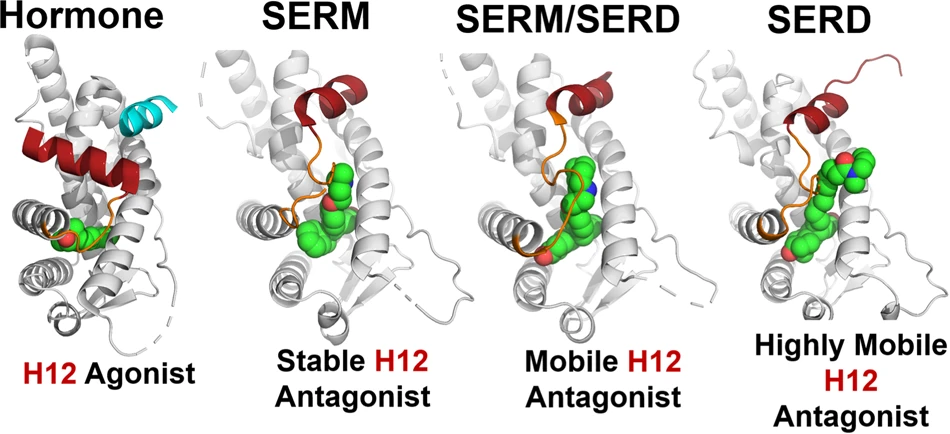
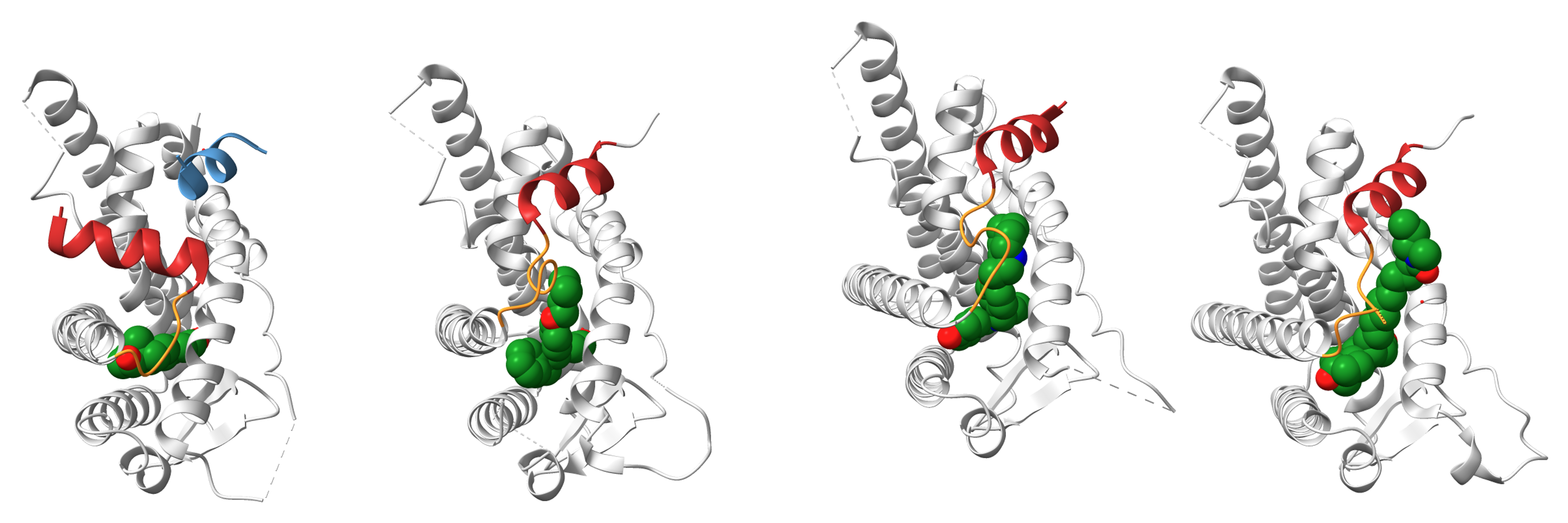
9.5 Tiled views
ChimeraX also supports tiled layouts, allowing multiple structures to be displayed side-by-side. This is particularly useful when comparing:
- different ligand-bound structures
- conformational states
- homologous proteins
The tile command automatically arranges open models into a grid layout. For example:
tile column 4 spacingfactor 0.8This arranges four models in a single row, with slightly reduced spacing between panels. Tiled views are frequently used in publications to compare related structures under consistent orientations.